Sometimes, the solution space in search and optimization problems cannot be expressed as a tree / graph of options (see previous AI post) or it cannot be easily traversed by heuristics or greedy algorithms. In those situations, we often use stochastic methods, those that heavily rely on a well described and rational random distribution. Genetic algorithms (or its broader concept of genetic programming) are one of such techniques.
Genetic algorithms are fascinating although they might look awkward at first. Do they really work? This is a question I have been asked many times. In fact, they do!
Genetic algorithms mimic the natural laws of evolution of living organisms that use genes as a way to code a solution to the problem of surviving in a specific environment. Such natural laws rely on natural selection and reproduction in a species to generate a population of best fit individuals.
For the sake of simplicity and clarity, we will use an example we described in our previous posts. Let’s imagine we want to design a molecule that best resembles a molecular reference by joining together 4 molecular building blocks from a pool of 65,536 building blocks. We will assume that any combination of building blocks is possible, that building blocks can be repeated, that the order of building blocks matters, and we will not take into consideration the chemical reactions required to link them.
Although the field has been widely studied and there are many aspects of genetic algorithms that can be solved differently (how to initialize the first generation or population, how to select individuals to cross over, how to mutate the individuals, how to measure the quality of the individuals, when to finish the search, …), we will use a few simple methods to understand the concept. In addition, most of these different methods depend on the particularity of the problem and they often need to be defined by human expertise.
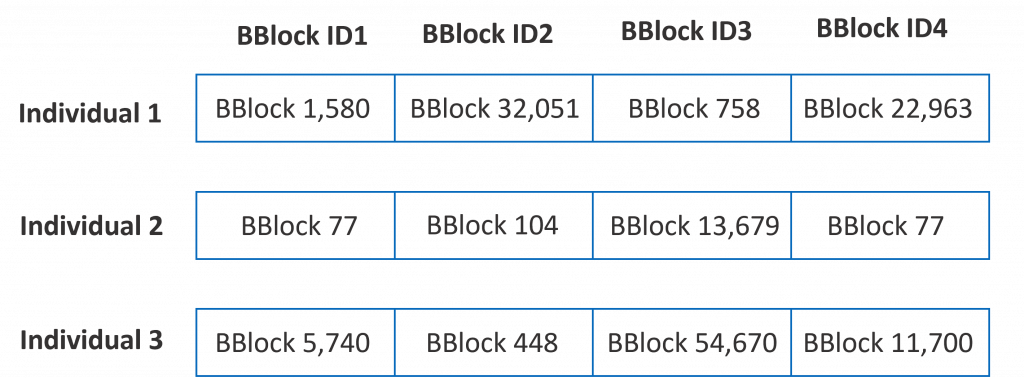
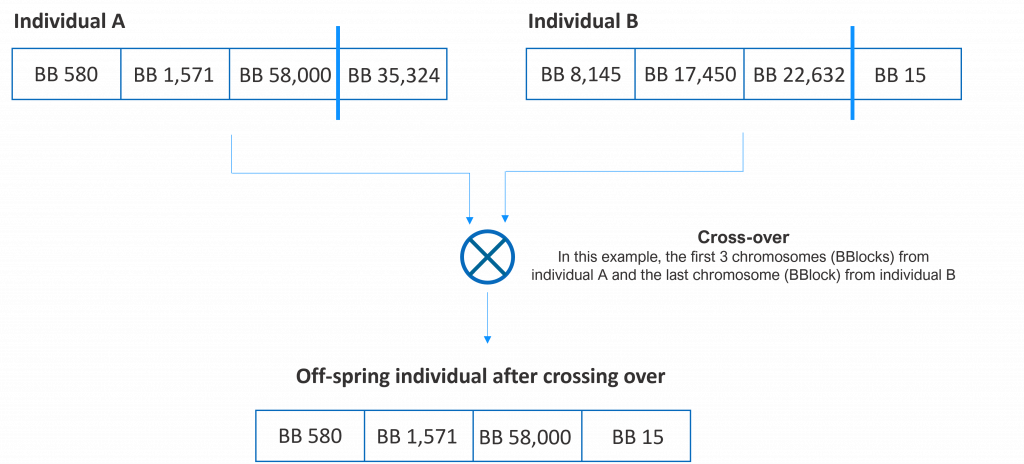
In a genetic algorithm, an individual (or a solution to a problem) is digitally coded as a set of chromosomes, each chromosome representing a specific characteristic of the individual. In our particular case, each of the 4 building blocks of a potential molecule is a chromosome. Hence, a valid molecule (a valid solution to the problem of finding a molecule similar to a reference molecule) is coded as an array of 4 16-bit identifiers (we need 16 bits to code one of the 65,536 possible building blocks). Find below 3 examples of possible individuals (solutions to the problem).

We also need to define a fitness function, which describes how well the individual (the solution) satisfies the problem. In our case, we can use a Tanimoto-like metric to compare each molecule (each solution) to the reference molecule. We want to maximize this value.
Initially, the algorithm randomly generates a population of individuals (solutions). Let’s say a population of 10,000 randomly generated molecules (the identifiers of the building blocks are set to valid but arbitrary values). This population will become the first generation of the genetic algorithm and it will probably contain very poor individuals since their Tanimoto values will be very low (expected from a random population).
Next, the algorithm iterates over several generations in order to improve the individuals and to enhance the solution to the problem. At each iteration or generation, the algorithm selects individuals of the current population and it performs two different actions on them (cross over and mutation) to generate a similar size population (next generation) that will replace the previous one. The amount of iterations is very dependent on the problem. We can fix a number, we can define a convergence metric of the population or a combination of both (just in case the problem does not converge or it takes too long to do so). Let’s say we will iterate for 5,000 generations.
As commented, at each iteration, the algorithm selects pairs of two individuals, it crosses them over (reproduce) to generate a new individual, it might mutate it and it adds the new individual to the new generation (population). The iteration will be finished once the number of individuals in the next generation is equal to the number of individuals of the current generation.
In this select+cross+mutate cycle, the selection function prioritizes individuals from the current population that have a better fitness score. In this sense, it mimics natural selection by enabling good individuals (best fit) to have higher chances to generate off-spring (new solutions). The selection function also chooses poor individuals, although with less probability, to have a more diverse search space.
Crossing over (reproducing) two individuals means combining their chromosomes and generating a new individual (off-spring). Although several options exist, in our example, we will pick a random number of chromosomes from the first individual of the pair and the rest from the second. In the following figure, we show several potential offspring of two individuals.
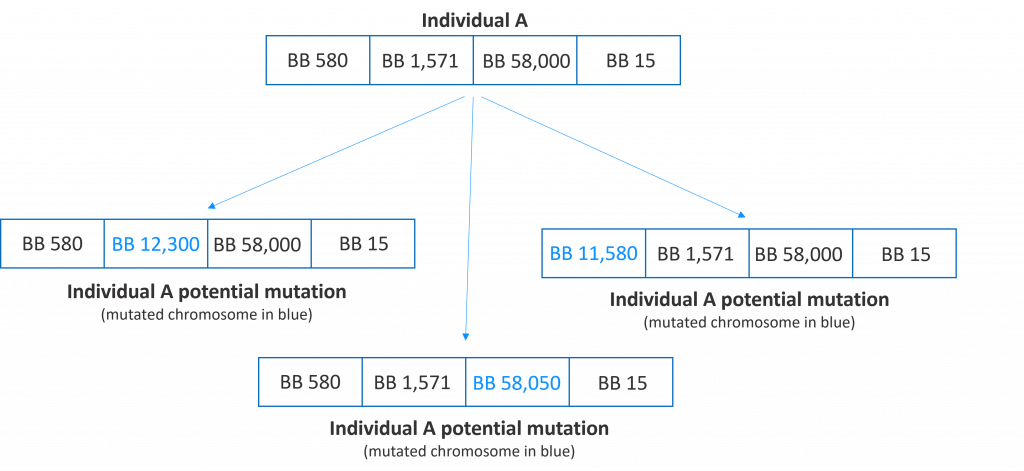
Mutating an individual means randomly changing one or more of its chromosomes. Mutation is an action that needs to occur rarely but it will allow the algorithm to jump to a new part of the search space. In the following figure, we show several potential mutations of an individual.
At the end of all iterations, the best fit or the set of best fit individuals are the solutions that the algorithm has found.
The following pseudo-code depicts the behavior of the described algorithm. As a stochastic method, genetic algorithms are not deterministic in nature and will generate different solutions each time they are executed (there are mechanisms to make them “randomly deterministic” for debugging or reproducibility purposes).
Initialize random population P of X individuals
For a number of iterations do
Create an empty new population P’
Repeat until new population P’ has reached X individuals
Select 2 individuals from current population P
Cross over these 2 individuals to create a new individual
Potentially mutate the chromosomes of the new individual with a low probability
Add the new individual to the new population P’
End repeat
Make the new population P’ be the current population P
End for
Select the best individual/s as the pseudo-optimal solution/s
Graphically, the evolution of the population and the search for a solution would look like the following figures.
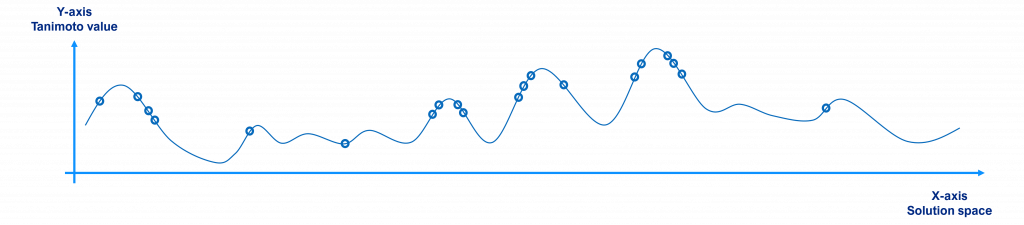
Initially, a randomly selected population will have individuals scattered around the search space as shown in the figure below, where the x-axis represents the solution space (each point being a different molecule or solution to the problem), the y-axis their Tanimoto value with respect to the reference (the value to maximize) and the circles the individuals of the population (for the sake of clarity we have just plotted 20 individuals, not 10,000).
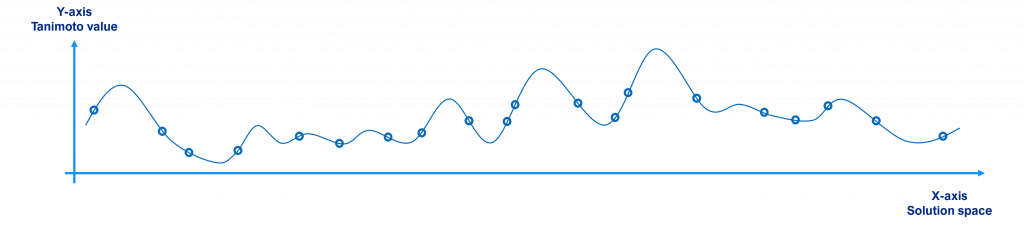
Over time, since better individuals have more chances to be selected and crossed over, the population tends to the local / global maxima points, as shown in the figure below.
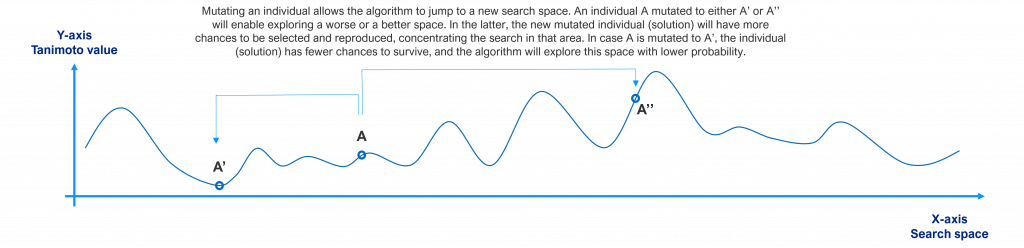
When the algorithm chooses to mutate an individual, it jumps to a new search space. If that new space is better than the current ones, this individual will have more probability to be selected and crossed over and the population will tend to move there. If the new space is worse, it does not have high chances to be selected and crossed over (low chances to survive). This is shown in the following picture for a single individual.
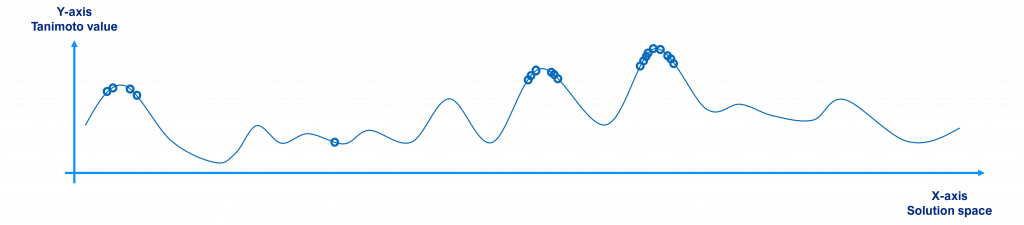
At the end, once the algorithm has iterated several times, the best individual or the best diverse individuals of the last population are the solutions found by the algorithm, as shown below. As commented before, the individuals of the last generation tend to be located near the local / global maxima points.
I hope this post has raised your interest in genetic algorithms and genetic programming and you will be eager to further read on this topic. Stay tuned for future Artificial Intelligence and Artificial Intelligence for Drug Discovery posts.
Want to learn more about how Pharmacelera can help you? Contact us and a member of our team will contact you to evaluate your project and how we can assist you with our technologies and services.