Fernando Martin
Pre & Post Sales Manager
The technology behind a software is important, but so is how to access it. The user interface (UI) allows users to effectively interact with the software and take advantage of the technology behind it. That’s why it’s important to provide users with user-friendly, powerful and intuitive UIs.
Each user is unique and has different needs, both in terms of preferences and the resources available to them. It is therefore important to offer different methods to suit those needs. In this article we will discuss the different ways to access Pharmacelera technologies.
Command-line Interface (CLI)
The CLI is probably one of the most widespread ways to interact with PharmScreen, exaScreen and PharmQSAR. This is the preferred option when using Linux operating systems through terminal applications, having access to all software functionalities. For other operating systems, you can take advantage of the Application Programming Interface (see below). Some of the features of the CLI are:I
- It is a versatile way to interact PharmScreen, exaScreen and PharmQSAR with the system environment, having good control over the execution process.
- You can leverage programming languages such as Bash or Python to automate certain processes (e.g., running several virtual screening campaigns at once).
- You need fewer hardware resources: Graphical environments require more hardware resources for molecular visualization.
- It allows you to connect to external resources such as other workstations, high performance computing (HPC) clusters, where there are usually no graphical interfaces.
- You can run graphical applications, such as MolXplore, to analyze screening results.
Application Programming Interface (API)
The API is another way to interact with our software. The API will allow you to run our tools on any OS, thanks to its integration with Docker. Some of the API features are:
- The ability to run Pharmacelera technologies on any operating system.
- To use simple configuration files to run your screening campaigns.
- To run your computations on external workstations, HPC clusters or cloud services, such as Amazon Web Services (AWS).
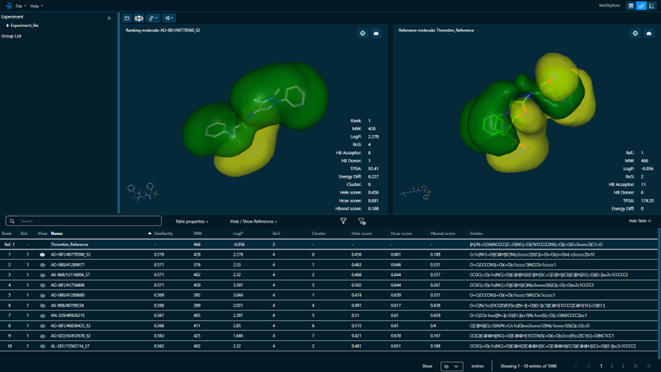
MolXplore
MolXplore is our GUI for analyzing the results obtained by our technologies. This tool, compatible with any operating system or accessible via web browser, provides a modern and intuitive environment to have a deeper understanding of your screening results, helping you to select the right compounds for later phases.
You can find the features and options of MolXplore in this post.
Knime nodes
The Knime platform is a graphical solution to build workflows that can help you analyze your data. The Pharmacelera extension (PharmScreen and exaScreen nodes) can easily connect to chemoinformatics nodes to run and analyze your virtual screening campaigns, connecting the results to other nodes for further analysis.
Some of the features of the Pharmacelera nodes developed for Knime are:
- An intuitive graphical user interface for setting up and running a virtual screening campaign using PharmScreen or exaScreen.
- The nodes use the Pharmacelera API, which means you can use Knime nodes on any operating system and run your screenings locally or on a remote cluster.
- You can reuse existing workflows created by Pharmacelera or the Knime community, making it easier to analyze your data.

Want to learn more?
If you would like to evaluate how our software can help you in your drug discovery projects or are looking for more customized services, please do not hesitate to contact us. We will provide you with personalized assistance so that you can find the best solution for your research.
